Awesome
CIAtah
<!-- # `CIAtah` (calciumImagingAnalysis [ciapkg]) -->
<!-- <img src="https://visitor-badge.glitch.me/badge?page_id=bahanonu.calciumImagingAnalysis" onerror="this.style.display='none'" alt='visitors' style='display:inline'> -->
<!--  -->
<!-- <img src="https://hitcounter.pythonanywhere.com/count/tag.svg?url=https%3A%2F%2Fgithub.com%2Fbahanonu%2FcalciumImagingAnalysis" onerror="this.style.display='none'" alt=''> -->
<img src="https://user-images.githubusercontent.com/5241605/117930569-03593480-b2b3-11eb-87f2-314e8ed77e94.png" align="center" onerror="this.style.display='none'" alt=''>
<br>
CIAtah (pronounced cheetah; formerly <ins>c</ins>alcium<ins>I</ins>maging<ins>A</ins>nalysis [ciapkg]) is a software package for analyzing one- and two-photon calcium imaging datasets. It can also be used to process other imaging datasets (e.g. from non-calcium indicators and dyes).
CIAtah currently requires MATLAB and runs on all major operating systems (Windows, Linux [e.g. Ubuntu], and macOS).
- Note:
CIAtahversionv4moves the remaining (i.e. all except external packages and software) CIAtah functions into theciapkgpackage to improve namespace handling and requires MATLAB R2019b or above (due to package import changes).
Full documentation at https://bahanonu.github.io/ciatah/.
Below are recordings and additional documents for users who want to learn more about calcium imaging analysis/experiments and the CIAtah pipeline.
<ins> Book chapter </ins> — We have a book chapter that goes over all steps of miniscope imaging: viral injections, GRIN lens probe implant, pain experimental design, data processing and neural/behavioral analysis, and more.
- See Ahanonu, B., Corder, G. (2022). Recording Pain-Related Brain Activity in Behaving Animals Using Calcium Imaging and Miniature Microscopes (https://doi.org/10.1007/978-1-0716-2039-7_13).
<ins>Webinar</ins> — This webinar gives an overview of calcium imaging analysis (with a focus on CIAtah) along with tips for improving experiments and analysis: https://info.inscopix.com/inscopix-inspire-view-webinarbiafra-ahanonu-signal-in-the-noise-distinguishing-relevant-neural-activity-in-calcium-imaging.
<ins>Workshop tutorial</ins> — This recording gives an overview of setting up and using CIAtah: https://www.youtube.com/watch?v=I6abW3uuJJw.
<ins>Imaging analysis tools</ins> My table with many current imaging analysis tools: https://github.com/bahanonu/imaging_tools.
<ins>GRINjector</ins> — A surgical device to help with implanting gradient-refractive index (GRIN) lens probes into the brain or other regions: https://github.com/bahanonu/GRINjector.
Spinal cord imaging
<ins>New motion correction methods</ins> — Methods for motion correction of spinal imaging data using feature identification (e.g. with DeepLabCut), control point registration, and other methods. Additional updates on integration into CIAtah in the future.
- Publication: B. Ahanonu*, A. Crowther*, A. Kania, M. Rosa-Casillas, A.I. Basbaum. (2024). Long-term optical imaging of the spinal cord in awake behaving mice. Nature Methods. https://doi.org/10.1038/s41592-024-02476-3 (https://www.nature.com/articles/s41592-024-02476-3)
- Preprint: Ahanonu and Crowther, et al. (2023). Long-term optical imaging of the spinal cord in awake, behaving animals. bioRxiv (https://www.biorxiv.org/content/10.1101/2023.05.22.541477v1.full).
- Documentation for spinal cord motion correction can be found at https://bahanonu.github.io/ciatah/pipeline_detailed_spinal.
- Other documentation for spinal cord imaging at https://github.com/basbaumlab/spinal_cord_imaging.
Contents
- CIAtah features
- CIAtah example features
- Quick start guide
- Quick start (command-line)
CIAtahmain GUI notes- Acknowledgments
- References
- Questions
- License
Contact: Biafra Ahanonu, PhD (github [at] bahanonu [dot] com).
Made in USA.<br> <img src="https://user-images.githubusercontent.com/5241605/71493809-322a5400-27ff-11ea-9b2d-52ff20b5f332.png" align="center" title="USA" alt="USA" width="auto" height="50">
CIAtah features
CIAtahpackage-enclosed functions (in+ciapkgsub-folders) can be used to create GUI-less, command line-ready analysis pipelines. As all functions are within theciapkgpackage for improve namespace handling to allow incorporating into other programs.- A GUI, via
ciatahclass, with different modules for large-scale batch analysis. - Includes all major calcium imaging analysis steps:
- movie visualization (including reading from disk, for fast viewing of large movies);
- pre-processing (motion correction [e.g. TurboReg, NoRMCorre] , spatiotemporal downsampling, spatial filtering, relative fluorescence calculation, etc.);
- support for multiple cell-extraction methods:
- <a href='https://github.com/mukamel-lab/CellSort'>PCA-ICA</a>
- <a href='https://searchworks.stanford.edu/view/11513617'>CELLMax</a> (<a href='https://searchworks.stanford.edu/view/12854822'>additional</a>)
- <a href='https://github.com/flatironinstitute/CaImAn-MATLAB' target='_blank'>CNMF</a>
- <a href='https://github.com/zhoupc/CNMF_E'>CNMF-E</a>
- <a href='https://github.com/schnitzer-lab/EXTRACT-public' target='_blank'>EXTRACT</a>
- etc.
- manual classification of cells via GUIs;
- automated cell classification (i.e. CLEAN algorithm, coming soon!);
- cross-session cell alignment;
- and more.
- Includes example one- and two-photon calcium imaging datasets for testing
CIAtah. - Supports a plethora of major imaging movie file formats: HDF5, NWB, AVI, MP4, ISXD [Inscopix], TIFF, BigTIFF, SLD [SlideBook], and Bio-Formats compatible formats (Olympus [OIR] and Zeiss [CZI and LSM] currently, additional support to be added or upon request).
- Supports Neurodata Without Borders data standard (see calcium imaging tutorial) for reading/writing cell-extraction and imaging movie files.
- Animal position tracking (e.g. in open-field assay) via ImageJ plugin.
- Requires
MATLABand runs on all major operating systems (Windows, Linux [e.g. Ubuntu], and macOS).
CIAtah example features
<p align="center"> <strong>Support for entire calcium imaging pipeline.</strong> </p> <!-- 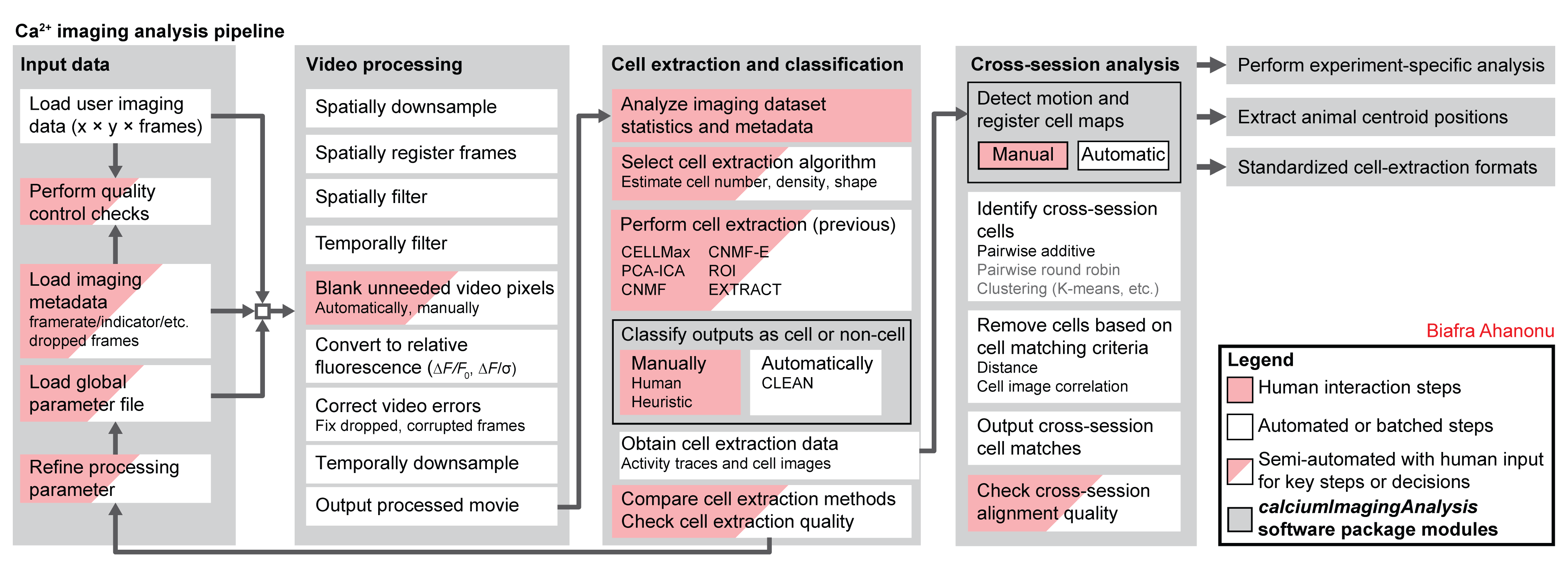 --> <!-- 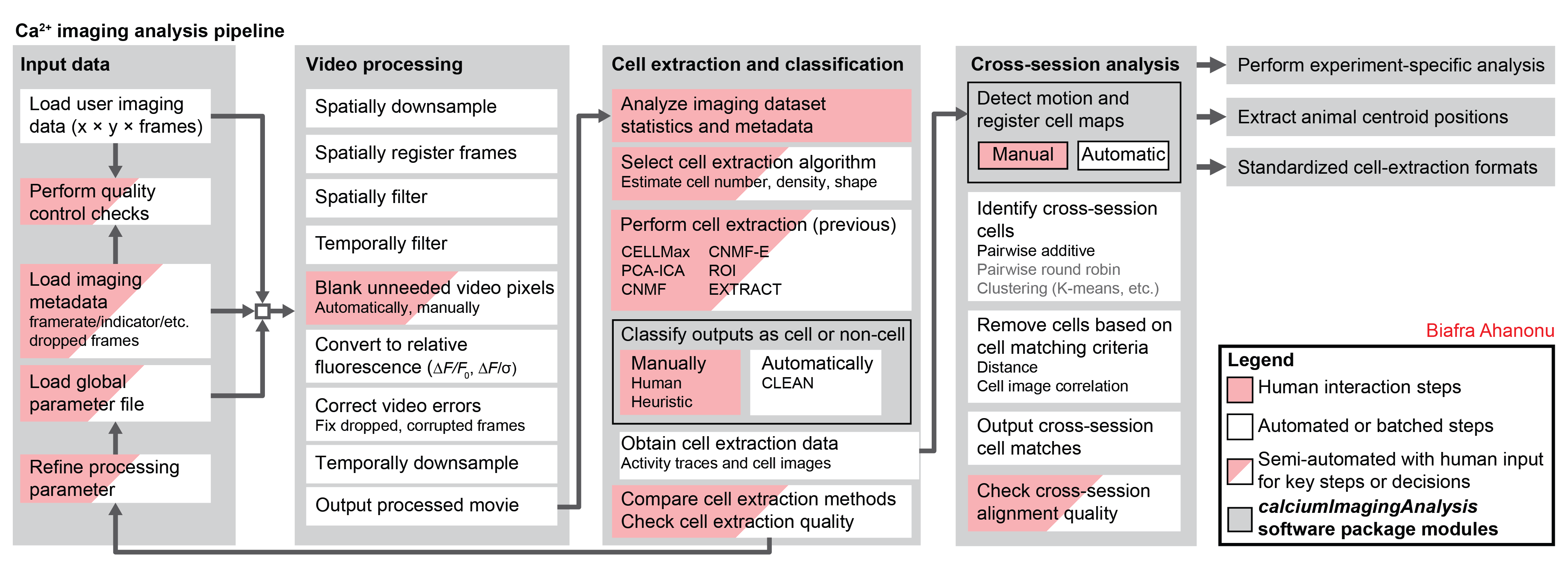 --> <p align="center"> <a href="https://github.com/bahanonu/ciatah/assets/5241605/9236ec16-38a9-40da-8545-ad40d0ea83c3"> <img src="https://github.com/bahanonu/ciatah/assets/5241605/9236ec16-38a9-40da-8545-ad40d0ea83c3" align="center" title="ciapkgMovie" alt="ciapkgMovie" width="100%" style="margin-left:auto;margin-right:auto;display:block;margin-bottom: 1%;"> </a> </p> <p align="center"> <strong>Movie processing, cell extraction, and analysis validation.</strong> </p> <p align="center"> • Press play if auto-play disabled. </p> <p align="center"> <a href="https://user-images.githubusercontent.com/5241605/94530890-9c3db280-01f0-11eb-99f0-e977f5edb304.gif"> <img src="https://user-images.githubusercontent.com/5241605/94530890-9c3db280-01f0-11eb-99f0-e977f5edb304.gif" align="center" title="ciapkgMovie" alt="ciapkgMovie" width="60%" style="margin-left:auto;margin-right:auto;display:block;margin-bottom: 1%;"> </a> </p> <p align="center"> <strong>Cell sorting GUI.</strong> </p> <p align="center"> • Press play if auto-play disabled. </p> <p align="center"> <a href="https://user-images.githubusercontent.com/5241605/100851700-64dec280-343a-11eb-974c-d6d29faf9eb2.gif"> <img src="https://user-images.githubusercontent.com/5241605/100851700-64dec280-343a-11eb-974c-d6d29faf9eb2.gif" align="center" title="ciapkgMovie" alt="ciapkgMovie" width="60%" style="margin-left:auto;margin-right:auto;display:block;margin-bottom: 1%;"> </a> </p> <p align="center"> <strong>Stable cell alignment across imaging sessions.</strong> </p> <p align="center"> • Press play if auto-play disabled. </p> <p align="center"> <a href="https://user-images.githubusercontent.com/5241605/105437652-4ca74a80-5c16-11eb-893a-87ea6d53e964.gif"> <img src="https://user-images.githubusercontent.com/5241605/105437652-4ca74a80-5c16-11eb-893a-87ea6d53e964.gif" align="center" title="m121_matchedCells" alt="m121_matchedCells" width="20%" style="margin-left:auto;margin-right:auto;display:block;margin-bottom: 1%;"> </a> </p> <!-- <p align="center"> <strong>Stable cell alignment across imaging sessions.</strong> </p> <p align="center"> <a href="https://user-images.githubusercontent.com/5241605/105437652-4ca74a80-5c16-11eb-893a-87ea6d53e964.gif"> <img src="https://user-images.githubusercontent.com/5241605/105437652-4ca74a80-5c16-11eb-893a-87ea6d53e964.gif" align="center" title="m121_matchedCells" alt="m121_matchedCells" width="20%" style="margin-left:auto;margin-right:auto;display:block;margin-bottom: 1%;"> </a> </p> -->Quick start guide
Below are steps needed to quickly get started using the CIAtah software package in MATLAB.
Download and install CIAtah
- Clone the
CIAtahrepository (using GitHub desktop or command line) or download the repository zip and unzip (e.g. run below MATLAB command).- Point the MATLAB path to the
CIAtahroot folder (NOT@CIAtahsub-folder in the repository). - Alternatively (not recommended since lags GitHub repository), download the package from
File Exchangeusing the Add-Ons explorer in MATLAB. SeeCIAtahentry at:or https://www.mathworks.com/matlabcentral/fileexchange/75466-ciatah.
- Point the MATLAB path to the
% Optional: this will set MATLAB working folder to the default user path. Make sure you have read/write permissions.
try; cd(userpath); catch; end;
% Download and unzip current repository
unzip('https://github.com/bahanonu/ciatah/archive/master.zip');
% Make CIAtah the working folder
cd('ciatah-master')
Check required toolboxes are installed
CIAtah depends on several MATLAB toolboxes to run properly. Run the below command to have CIAtah check whether dependencies have been installed properly. If not, use the Add-Ons (https://www.mathworks.com/help/matlab/matlab_env/get-add-ons.html) explorer to install each toolbox.
ciapkg.io.matlabToolboxCheck();
Setup CIAtah
- Download and install
CIAtahdependencies. These will all be located in the_external_programssub-folder within the CIAtah main code directory.
ciapkg.io.loadDependencies;
- Run
CIAtahusing the below MATLAB commands. Callobj;in the MATLAB command window each time you want to go back to the main GUI.- Note:
calciumImagingAnalysisclass is now calledciatah, all functionality is the same.
- Note:
% Loads the class into an object for use in this session
obj = ciatah();
% Runs routines to check dependencies and help user get setup.
obj.setup();
% Open the class menu (always type `obj` then enter load the class/modules menu)
obj % then hit enter, no semicolon!
- Afterwards, run the
modelAddNewFoldersmodule to add data folders the currentCIAtahclass object. - Run
obj;in the command window to see the main GUI. - Full documentation at https://bahanonu.github.io/ciatah/.
- [Optional] Users on Windows systems can download
Everything(https://www.voidtools.com/). It is a very useful and extremely fast search engine for files and folders that allows users to quickly obtain lists of full folder paths for analysis inCIAtah. - [Optional] If run into issues opening certain AVI files (e.g. due to codec issues) with CIAtah/MATLAB, install
K-Lite Codec Pack(https://codecguide.com/download_kl.htm) or similar for added support.
Visualize movies quickly using read from disk
Method #1: CIAtah playMovie
Users can quickly visualize movies in any of the supported formats (HDF5, NWB, AVI, TIFF, ISXD, etc.) using the playMovie function. This will read directly from disk, allowing users to scroll through frames to visually check movies before or after processing.
Users can run via the command-line:
% Use the absolute path to the movie file or a valid relative path.
ciapkg.api.playMovie('ABSOLUTE_PATH_TO_MOVIE\MOVIE_NAME.EXTENSION');
When using HDF5 files, check the dataset name containing movie with h5disp then input the full dataset name (e.g. below is for a standard NWB-formatted HDF5 file):
ciapkg.api.playMovie('ABSOLUTE\PATH\TO\MOVIE','inputDatasetName','/acquisition/TwoPhotonSeries/data');
Method #2: CIAtah GUI
Using the ciatah GUI class, users can select loaded folders and change the regular expression to match the name of the files in the movie, both for the raw data and for any processed movies in the folder. See below:
Method #3: Fiji + N5
Install N5 library to enable N5 viewer in Fiji: go to Help->Update... then under Manage Update Sites scroll down to select N5 (https://sites.imagej.net/N5/) then Apply and Close). View a large HDF5 movie by running File->Import->"HDF5/N5/Zarr/OME-NGFF". In the resulting dialog box, select Browse along with Open as virtual on the bottom left. After a moment different a tree should appear listing different datasets
- https://imagej.net/libs/n5
- https://github.com/saalfeldlab/n5
- https://github.com/saalfeldlab/n5-ij
- https://github.com/saalfeldlab/n5-viewer
Quick start (command line or GUI-less batch analysis)
After downloading CIAtah and running the setup as above, users interested in command-line processing can open up the example M-file by running the below command. By running individual code-block cells, users are guided from pre-processing through cell-extraction to cross-session analysis.
edit ciapkg.demo.cmdLinePipeline
Users can import the CIAtah ciapkg that contains the command-line functions using the below command at the beginning of their functions. This will import all ciapkg functions into the functions workspace such that ciapkg.api.loadMovieList() (an alias for ciapkg.io.loadMovieList()) can be called as loadMovieList().
import ciapkg.api.* % import CIAtah functions in ciapkg package API.
CIAtah main GUI notes
See more about the CIAtah GUI at https://bahanonu.github.io/ciatah/install/#ciatah-main-gui-notes.
Acknowledgments
Thanks to Jones G. Parker, PhD (https://parker-laboratory.com/) for providing extensive user feedback during the development of the CIAtah software package.
Additional thanks to Drs. Jesse Marshall, Jérôme Lecoq, Tony H. Kim, Hakan Inan, Lacey Kitch, Maggie Larkin, Elizabeth Otto Hamel, Laurie Burns, and Claudia Schmuckermair for providing feedback, specific functions, or helping develop aspects of the code used in CIAtah.
References
Please cite Corder*, Ahanonu*, et al. 2019 Science publication or the Ahanonu, 2018 Zenodo release if you used the software package or code from this repository to advance/help your research:
@article{corderahanonu2019amygdalar,
title={An amygdalar neural ensemble that encodes the unpleasantness of pain},
author={Corder, Gregory and Ahanonu, Biafra and Grewe, Benjamin F and Wang, Dong and Schnitzer, Mark J and Scherrer, Gr{\'e}gory},
journal={Science},
volume={363},
number={6424},
pages={276--281},
year={2019},
publisher={American Association for the Advancement of Science}
}
Please cite the Ahanonu and Corder, 2022 book chapter if you used procedures detailed therein.
@incollection{ahanonu2022recording,
title={Recording Pain-Related Brain Activity in Behaving Animals Using Calcium Imaging and Miniature Microscopes},
author={Ahanonu, Biafra and Corder, Gregory},
booktitle={Contemporary Approaches to the Study of Pain},
pages={217--276},
year={2022},
publisher={Springer}
}
Please cite the Ahanonu*, Crowther*, 2024 Nature Methods paper on spinal cord imaging if you use related methods or procedures.
@article{Ahanonu2024-wq,
title = "Long-term optical imaging of the spinal cord in awake behaving
mice",
author = "Ahanonu, Biafra and Crowther, Andrew and Kania, Artur and
Rosa-Casillas, Mariela and Basbaum, Allan I",
journal = "Nat. Methods",
publisher = "Springer Science and Business Media LLC",
pages = "1--13",
month = nov,
year = 2024,
language = "en"
}
Please see https://bahanonu.github.io/ciatah/references/ for additional references depending on processing steps undertaken.
<!-- ```Latex @misc{biafra_ahanonu_2018_2222295, author = {Biafra Ahanonu}, title = {{calciumImagingAnalysis: a software package for analyzing one- and two-photon calcium imaging datasets.}}, month = December, year = 2018, doi = {10.5281/zenodo.2222295}, url = {https://doi.org/10.5281/zenodo.2222295} } ``` -->Questions?
Please email any additional questions not covered in the repository to github [at] bahanonu [dot] com or open an issue.
Users with versions of MATLAB earlier than R2019b can download CIAtah version v3 (see Releases) until pre-R2019b MATLAB support is fully integrated into v4.
License
Copyright (C) 2013-2024 Biafra Ahanonu
This project is licensed under the terms of the MIT license. See LICENSE file for details.
Repository stats
Statistics on total hits on the main repository and documentation pages. Several badges are present as certain services have gone offline or count page hits differently.
<!-- - [](http://hits.dwyl.com/bahanonu/calciumImagingAnalysis) (starting 2020.08.16), frozen til `dwyl` migrates to new server. -->



